Resources: Posters
Integrated Morphology-Transcriptomics Analysis Reveals Functional Cell States in Macrophage Polarization and Tumor-Induced Reprogramming
February 23, 2026
Keystone | Myeloid Cells: Functional Heterogeneity with Therapeutic Promise
Shreya Deshukh Ph.D., Sr. Data Scientist
Keywords: Tumor microenvironment, Myeloid cells, Morphological profiling, Immune reprogramming, Oncology

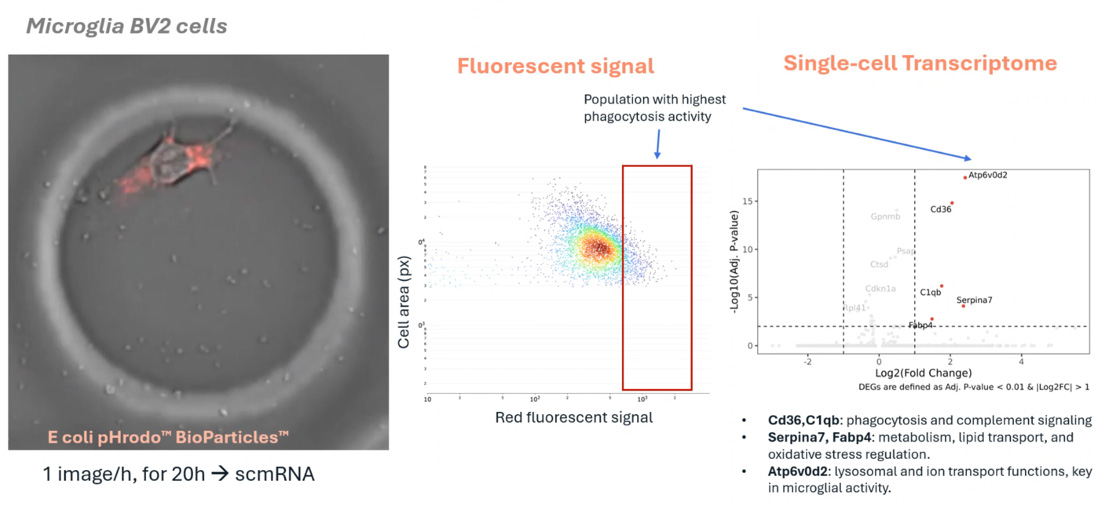
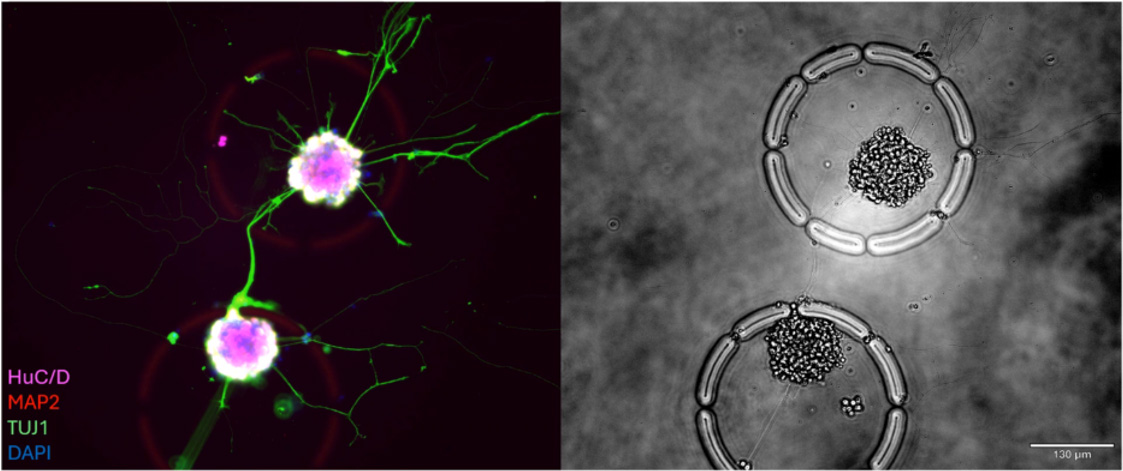
This research explores the relationship between structural changes and functional states during macrophage polarization and tumor-induced reprogramming. Using integrated morphology-transcriptomics, the platform identifies key indicators of immunosuppressive transitions and provides a high-resolution map of the tumor microenvironment.
FAQ's
How does the analysis correlate morphology with macrophage functional states?
Macrophages undergo significant structural transitions during polarization. This integrated analysis maps specific morphological signatures to transcriptomic profiles, revealing how tumor-induced reprogramming alters both the physical characteristics and the functional immune response of the cells.
Can this system identify tumor-induced reprogramming in single cells?
The system tracks macrophage transitions in response to tumor environments in real time. By linking live-cell behavior to RNA programs, the analysis pinpoint the triggers of immunosuppressive reprogramming at single-cell resolution.