Resources: Posters
Novel CellCage™ technology integrates image-based phenotyping and single-cell transcriptomics to study dynamic behaviors of living cells
December 1, 2025
ASCB | American Society of Cell Biology
Frank Charbonier, Ph.D., Biomaterials Scientist
Keywords: Image-based phenotyping, Single-cell transcriptomics, Multi-modal analysis, Live-cell dynamics, R3200 Platform, CellCage™ technology

Leveraging the R3200 Platform and its foundational CellCage™ technology, this research demonstrates the integration of biocompatible, light-guided hydrogel microenvironments to bridge the gap between live-cell imaging and high-resolution transcriptomics. By securing individual cells or defined groups in configurable enclosures, the platform enables the long-term study of phagocytosis, cell-cell interactions, and calcium signaling. This integrated approach allows for the direct mapping of complex, image-based phenotypes to underlying genetic programs, providing a scalable alternative to traditional droplet or nanowell methods
FAQ's
How does CellCage™ technology on the R3200 Platform differ from standard nanowell or droplet-based methods?
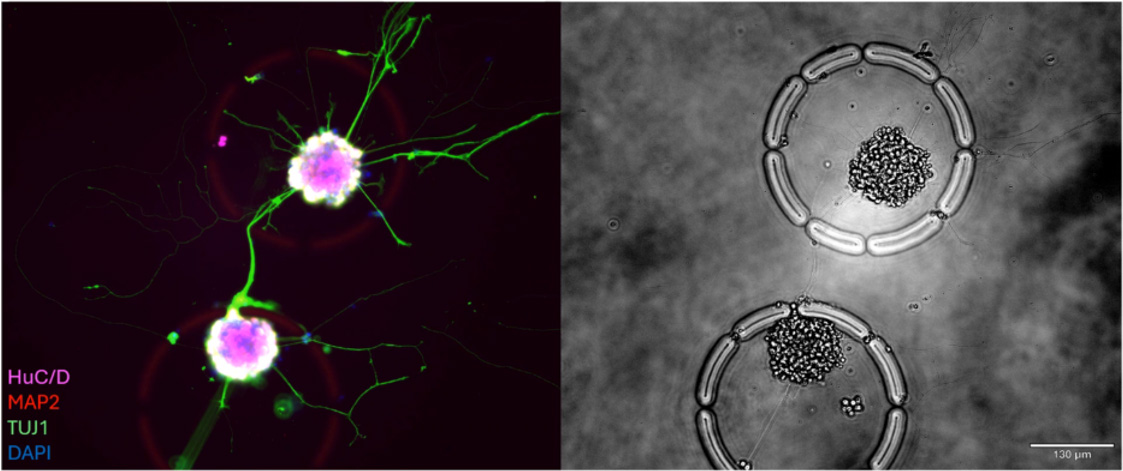
Unlike static well-plates or droplets, CellCage™ enclosures (CCEs) are biocompatible hydrogel microenvironments that can be configured in size, shape, and porosity. This allows the R3200 Platform to support the longitudinal culture of suspension or adherent cells for weeks while maintaining permeable walls for nutrient exchange and real-time monitoring.
How is the R3200 Platform used to link dynamic behaviors to transcriptomic data?
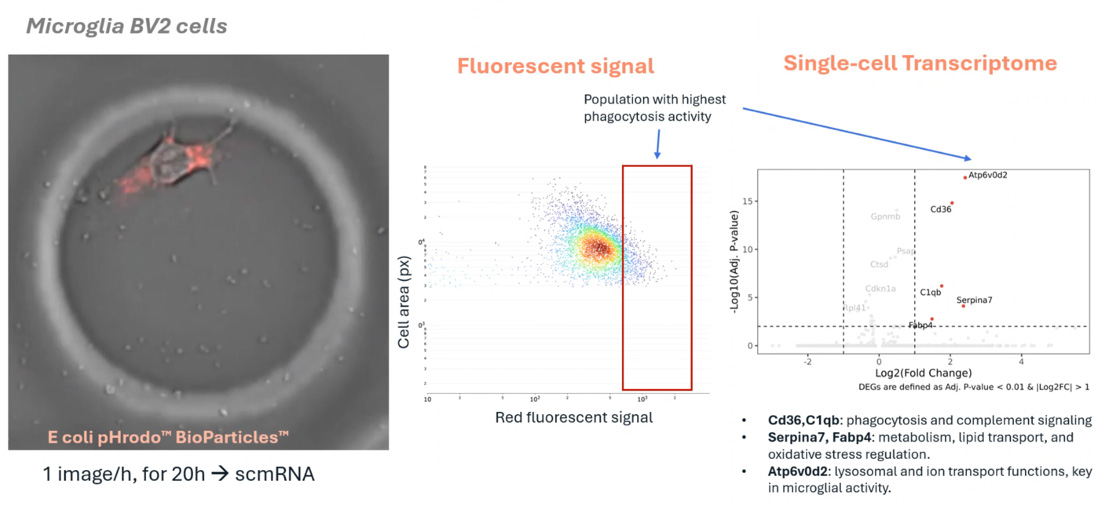
The R3200 Platform facilitates continuous, high-resolution imaging of dynamic behaviors—such as microglial phagocytosis or T-cell activation—while cells are secured within CCEs. Following the observation period, the platform performs direct transcriptomic profiling on those same individual cells, providing a definitive link between a cell's observed history and its molecular state.
What emerging capabilities does the R3200 Platform enable through selective degradation?
Utilizing the R3200's ability to selectively degrade CellCage™ enclosures with cell-compatible chemistry, the technology enables the enrichment or retention of specific target cells from mixed populations. This allows for focused functional profiling and transcriptomic analysis of rare subsets, such as NK cells from unlabeled PBMC mixes, within a single integrated workflow.