
CRISPR Screens:
See Which Edits Change What Cells Do.
Function is observed directly, not inferred. Nominate hits you might otherwise miss.
The readout shapes the hit list.
- Many pooled screens rely on proxy readouts to infer the phenotype you care about.
- But a proxy is not just an imperfect measurement. It is a filter. It determines which perturbations can rise to the top. When the proxy and phenotype align, those screens can identify strong candidates. When they diverge, important perturbations may be deprioritized, while proxy-supported calls may move forward without enough functional evidence.
- Cellanome CRISPR Screening measures live-cell phenotype directly, then links those outcomes to morphology, guide identity, and the same-cell transcriptome to help explain why each perturbation matters.
Because RNA is not the only way cells change. Perturbations can alter morphology, protein regulation, timing, localization, post-translational modification, or cell-cell behavior without producing a proportional transcriptomic signature.
If that is the biology your screen is meant to find, the readout has to measure it directly.
Illustrative Polyfunctional Potency Screen
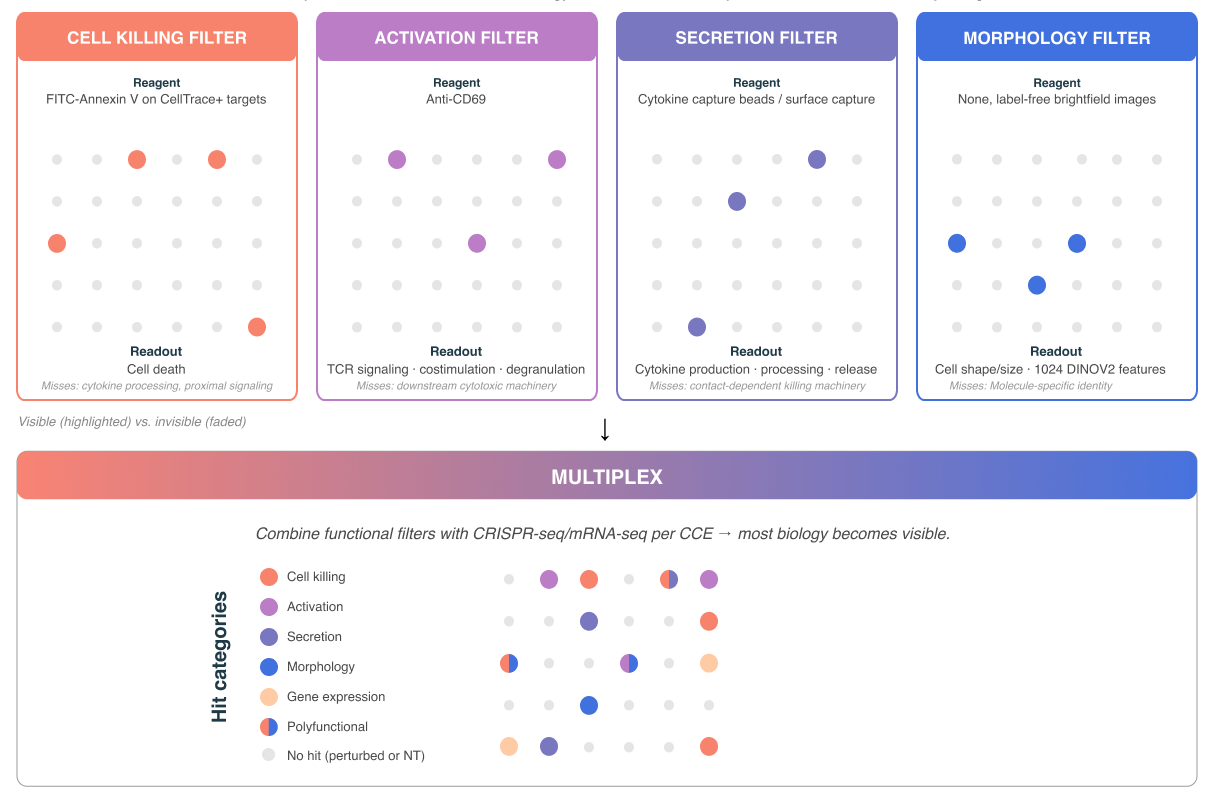
When this approach fits
Offered as a Cellanome-run service in our lab or as a supported workflow on the R3200 platform. Cellanome CRISPR screening is ideal for phenotypes that are dynamic, contextual, adherent, or multidimensional, especially when a single endpoint cannot fully address the biological question.
- Dynamic function over time. Secretion, proliferation, phagocytosis, differentiation, senescence, persistence, and recovery.
Phenotypes that emerge as cells behave over time, not when they are captured in a single snapshot. - Cell-cell interactions. Activation, killing, priming, pruning, signaling, and immune suppression.
Extended phenotypes that resolve when you measure how a perturbed cell changes its interaction with another. - Engineering trade-offs. CAR-T potency versus persistence, cell-line secretion versus proliferation, iPSC differentiation efficiency versus purity.
Screens where optimizing one axis without measuring the other lets variants escape on the unmeasured axis. - Same-cell multimodal readouts. Senescence, CAR-T polyfunctional potency, macrophage polarization, and cell-state transitions.
Phenotypes that require several measurements to remain linked to the same cell. - Adherent cells in their attached state. Neurons, fibroblasts, solid tumor cells, stromal cells, and glia.
Phenotypes that require cells to stay attached, viable, and morphologically intact.
Cellanome refines and expands CRISPR hit discovery
Refines: Distinguishes transcriptional hits with functional support from those without.
Expands: Surfaces functional and morphological hits that RNA-only screens can miss.
Because perturbation, behavior, morphology, and transcriptome remain linked, each hit can be ranked by evidence across biological layers.
Classify hits by evidence across biological layers

A Cellanome screen produces a sharper hit list:
- Fewer proxy-supported calls entering validation without functional evidence
- More functionally-relevant candidates entering validation
- Clearer links from perturbation to phenotype, mechanism, and downstream liabilities
- Perturbation- and function-labeled transcriptomes for every cell, including hits and non-hits, ready for hit classification, mechanism mapping, and model training
Examples of Cellanome CRISPR screening
Cell-Cell Interactions: Spitzer Lab, UCSF
The Spitzer Lab is applying this approach to immune-cell activation, where the phenotype of interest, T-cell priming, is not fully captured by any single surface marker.
Existing methods can measure proxies such as CD80 expression. Cellanome can measure the extended phenotype directly: how edits in one cell affect its interaction with another.
Proxy Readout

Intended Phenotype

Linking live-cell behavior to transcriptional responses across perturbations using dynamic caging
To demonstrate this approach, we developed Perturb-LINK (PERTURBational LINKage of transcriptomics and imaging in single cells via enclosure-based screening), a method that links guide identity with same-cell transcriptome, morphology, and surface protein readouts in a pooled screen.
Perturb-LINK applies the same phenotype-first logic to pooled CRISPR screening. In murine bone marrow-derived macrophages under LPS stimulation, guide identity was linked with same-cell transcriptome, morphology, and surface protein readouts.
Three illustrative perturbations landed in different layers of the multi-modal hit list:

A transcriptome-only screen would have captured only part of this biology. Cellanome helps identify which perturbations are functionally supported, which are RNA-silent but phenotypic, and which are best used for triage or model training.
Run your CRISPR screen with Cellanome. Or run it on your R3200.
Cellanome’s service lab can run the screen end-to-end. Or you can purchase the platform and run it in your own lab.
Most CRISPR engagements start by establishing your cell model and readouts on the R3200. Our field scientists can support your team directly, or our concierge service team can build and run the workflow.
Phase 1: Build the Workflow
Set up your cell model on the R3200. Together, we define what to measure, when, and how live-cell readouts, fluorescent channels, induction steps, and end-point sequencing fit together.
Output: A locked, multi-day workflow tuned to your cell model, with example data before the full screen begins.
Illustrative Polyfunctional Potency Screen Run Design: Multi-day Multimodal Workflow


Phase 2: Run the Screen
Run a targeted or genome-wide CRISPR screen. Review data as the screen progresses, with checkpoints along the way to adjust target coverage as signal emerges.
Output: A decision-support map that links each guide to observed phenotype, morphology, and same-cell transcriptome, with analysis-ready outputs for hit classification, validation planning, and model training.
Optional deeper analysis packages rank perturbations by evidence class, connect phenotypic hits to transcriptomic programs, and surface candidate mechanisms, potential liabilities, and model-ready phenotypic labels.
Talk to us about your screen
Tell us more about what you want to measure. We'll follow up to discuss the next steps for your lab.
